BLASTN 2.2.24 [Aug-08-2010]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= DPGLEAN22038 (UTR included)
(1594 letters)
Database: est
9484 sequences; 8,280,457 total letters
Searching..................................................done
Score E
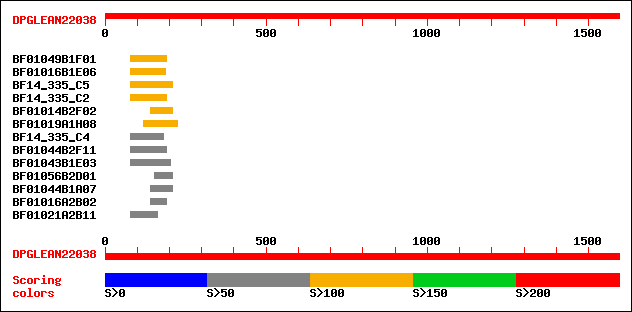
Sequences producing significant alignments: (bits) Value
BF01049B1F01 137 6e-32
BF01016B1E06 119 1e-26
BF14_335_C5 111 3e-24
BF14_335_C2 109 1e-23
BF01014B2F02 105 2e-22
BF01019A1H08 105 2e-22
BF14_335_C4 90 1e-17
BF01044B2F11 78 5e-14
BF01043B1E03 78 5e-14
BF01056B2D01 74 8e-13
BF01044B1A07 72 3e-12
BF01016A2B02 70 1e-11
BF01021A2B11 68 5e-11
>BF01049B1F01
Length = 840
Score = 137 bits (69), Expect = 6e-32
Identities = 102/113 (90%)
Strand = Plus / Minus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagatagggctcctcaaacc 138
||||||||||||||||| ||| || | |||||||| ||||||||||||| ||| | |||
Sbjct: 698 agaggagcttggacggtagaggtgggtgtttaacgggagttagatagggccccttacacc 639
Query: 139 aacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacga 191
|||||||||||||||||||||||| |||||||||||||||||||||||||||
Sbjct: 638 tacacctgggtcgcaagtcccaggcactgttgaagctcctctccctgcaacga 586
>BF01016B1E06
Length = 606
Score = 119 bits (60), Expect = 1e-26
Identities = 99/111 (89%), Gaps = 2/111 (1%)
Strand = Plus / Minus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagatagggctcctcaaacc 138
||||||| ||||||||||||| |||||||||||||| |||||||||||| ||| |||||
Sbjct: 301 agaggagattggacggtggaggtgagcgtttaacgc--gttagatagggccccttaaacc 244
Query: 139 aacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaac 189
||||||||||||| | |||| ||| ||||||||||||||||| |||||||
Sbjct: 243 tacacctgggtcgctactccccggcactgttgaagctcctctctctgcaac 193
>BF14_335_C5
Length = 1597
Score = 111 bits (56), Expect = 3e-24
Identities = 112/131 (85%), Gaps = 1/131 (0%)
Strand = Plus / Plus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagataggg-ctcctcaaac 137
||||||||||||||||||||| ||||||||||| || |||||| |||| | ||| ||||
Sbjct: 1446 agaggagcttggacggtggaggtgagcgtttaatgcgcgttagacaggggcgccttaaac 1505
Query: 138 caacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacgatggacc 197
| |||| ||||||||||||||| |||||||||||||||||| ||||| |||| |||
Sbjct: 1506 ctacacnnnnntcgcaagtcccaggcactgttgaagctcctctccgtgcaaagatgcacc 1565
Query: 198 cgccacccgca 208
|| ||||||||
Sbjct: 1566 cggcacccgca 1576
>BF14_335_C2
Length = 1827
Score = 109 bits (55), Expect = 1e-23
Identities = 100/114 (87%), Gaps = 2/114 (1%)
Strand = Plus / Minus
Query: 79 agaggagcttggacggtggagctgagcgttta--acgcaagttagatagggctcctcaaa 136
||||||||||||||||||||| |||||||| |||| |||||||||||| || |||
Sbjct: 976 agaggagcttggacggtggaggtgagcgttatgtacgcgagttagataggggctcttaaa 917
Query: 137 ccaacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacg 190
|| ||||||| ||||||| |||||||| ||||||||||||||||||||||||||
Sbjct: 916 cctacacctgtgtcgcaaatcccaggcactgttgaagctcctctccctgcaacg 863
>BF01014B2F02
Length = 623
Score = 105 bits (53), Expect = 2e-22
Identities = 65/69 (94%)
Strand = Plus / Minus
Query: 140 acacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacgatggacccg 199
||||||||||||||||||| |||||||| |||||||||||||||||||||| ||||||||
Sbjct: 622 acacctgggtcgcaagtcctaggcgctgctgaagctcctctccctgcaacgctggacccg 563
Query: 200 ccacccgca 208
||||||||
Sbjct: 562 gcacccgca 554
>BF01019A1H08
Length = 504
Score = 105 bits (53), Expect = 2e-22
Identities = 99/112 (88%), Gaps = 3/112 (2%)
Strand = Plus / Minus
Query: 118 ttagatagg-gctcctcaaaccaacacctgggtcgcaagtcccaggcgctgttgaagctc 176
||||||||| || ||| ||||| ||||||||||||||| |||||||| ||||||||||||
Sbjct: 492 ttagataggagccccttaaacctacacctgggtcgcaactcccaggcactgttgaagctc 433
Query: 177 ctctccctgcaa--cgatggacccgccacccgcatggccaatacattgaatg 226
|||||||||||| ||||||||||| |||||||| || ||| |||||||||
Sbjct: 432 ctctccctgcaacgcgatggacccggcacccgcacggtgaatccattgaatg 381
>BF14_335_C4
Length = 1491
Score = 89.7 bits (45), Expect = 1e-17
Identities = 91/105 (86%), Gaps = 1/105 (0%)
Strand = Plus / Plus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagatag-ggctcctcaaac 137
||||||||||||| || ||| ||||||||||| || || ||||| ||| ||| ||||
Sbjct: 1198 agaggagcttggaaagttgaggtgagcgtttaatgcgcgtaagataaaggccccttaaac 1257
Query: 138 caacacctgggtcgcaagtcccaggcgctgttgaagctcctctcc 182
| |||||||||||||||||||||||| ||||||||||||||||||
Sbjct: 1258 ctacacctgggtcgcaagtcccaggcactgttgaagctcctctcc 1302
>BF01044B2F11
Length = 728
Score = 77.8 bits (39), Expect = 5e-14
Identities = 97/114 (85%), Gaps = 3/114 (2%)
Strand = Plus / Plus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagatagggctc-ctcaaac 137
|||||||||||||| |||||| |||||||||| | || ||||||||| | || ||||
Sbjct: 598 agaggagcttggacagtggaggtgagcgtttatcacag--tagataggggccacttaaac 655
Query: 138 caacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacga 191
| ||||||| | |||||||||||| ||||| ||||||||||||||||||||||
Sbjct: 656 ctacacctgaattgcaagtcccaggtgctgtcgaagctcctctccctgcaacga 709
>BF01043B1E03
Length = 830
Score = 77.8 bits (39), Expect = 5e-14
Identities = 106/127 (83%), Gaps = 1/127 (0%)
Strand = Plus / Plus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagatagggctcctc-aaac 137
|||||| ||||||| |||||| ||||| ||||||| ||||||||||| ||| ||||
Sbjct: 342 agaggaccttggacagtggaggtgagcacttaacgcgcgttagataggggccctttaaac 401
Query: 138 caacacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacgatggacc 197
| | |||||||| |||||||||| || || ||| | |||||||||| |||||||||||||
Sbjct: 402 ctatacctgggttgcaagtcccaagcactattggacctcctctcccagcaacgatggacc 461
Query: 198 cgccacc 204
|| ||||
Sbjct: 462 cgtcacc 468
>BF01056B2D01
Length = 728
Score = 73.8 bits (37), Expect = 8e-13
Identities = 52/56 (92%), Gaps = 2/56 (3%)
Strand = Plus / Minus
Query: 155 gtcccaggcgctgttgaagctcctctccctgcaa--cgatggacccgccacccgca 208
|||||||||||||||||||||||||||||||||| |||||||||| ||||||||
Sbjct: 166 gtcccaggcgctgttgaagctcctctccctgcaatgtgatggacccggcacccgca 111
>BF01044B1A07
Length = 806
Score = 71.9 bits (36), Expect = 3e-12
Identities = 63/71 (88%), Gaps = 2/71 (2%)
Strand = Plus / Minus
Query: 140 acacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaa--cgatggacc 197
|||||||| |||| |||||||| | | |||||||||||||||||||| | |||||||||
Sbjct: 532 acacctggttcgcgagtcccagacacagttgaagctcctctccctgccacgcgatggacc 473
Query: 198 cgccacccgca 208
|||||||||||
Sbjct: 472 cgccacccgca 462
>BF01016A2B02
Length = 780
Score = 69.9 bits (35), Expect = 1e-11
Identities = 47/51 (92%)
Strand = Plus / Minus
Query: 140 acacctgggtcgcaagtcccaggcgctgttgaagctcctctccctgcaacg 190
||||||| ||||||| |||||||| |||||||||||||||||||| |||||
Sbjct: 274 acacctgagtcgcaattcccaggcactgttgaagctcctctcccttcaacg 224
>BF01021A2B11
Length = 729
Score = 67.9 bits (34), Expect = 5e-11
Identities = 74/86 (86%), Gaps = 1/86 (1%)
Strand = Plus / Plus
Query: 79 agaggagcttggacggtggagctgagcgtttaacgcaagttagat-agggctcctcaaac 137
||||||||||||||||||||| |||||||||||||| ||||||| |||| ||| |||
Sbjct: 375 agaggagcttggacggtggaggtgagcgtttaacgcgtgttagatcggggcgccttcaac 434
Query: 138 caacacctgggtcgcaagtcccaggc 163
| |||||||||| | ||||||||||
Sbjct: 435 ctacacctgggtgcccagtcccaggc 460
Database: est
Posted date: Jul 14, 2011 2:25 PM
Number of letters in database: 8,280,457
Number of sequences in database: 9484
Lambda K H
1.37 0.711 1.31
Gapped
Lambda K H
1.37 0.711 1.31
Matrix: blastn matrix:1 -3
Gap Penalties: Existence: 5, Extension: 2
Number of Sequences: 9484
Number of Hits to DB: 204,332
Number of extensions: 4608
Number of successful extensions: 4608
Number of sequences better than 1.0e-10: 13
Number of HSP's gapped: 4601
Number of HSP's successfully gapped: 13
Length of query: 1594
Length of database: 8,280,457
Length adjustment: 17
Effective length of query: 1577
Effective length of database: 8,119,229
Effective search space: 12804024133
Effective search space used: 12804024133
X1: 11 (21.8 bits)
X2: 15 (29.7 bits)
X3: 50 (99.1 bits)
S1: 12 (24.3 bits)
S2: 34 (67.9 bits)

