BLASTN 2.2.24 [Aug-08-2010]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= DPGLEAN19942 (UTR included)
(1747 letters)
Database: est
9484 sequences; 8,280,457 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF01031A2G02 88 6e-17
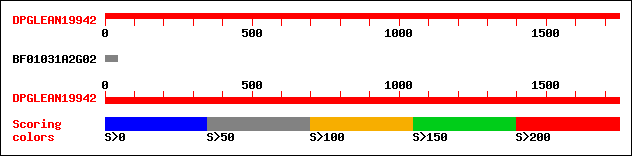
>BF01031A2G02
Length = 535
Score = 87.7 bits (44), Expect = 6e-17
Identities = 44/44 (100%)
Strand = Plus / Minus
Query: 1 caatattcgccatgtttaaagcgatactaaaatttccttcactt 44
||||||||||||||||||||||||||||||||||||||||||||
Sbjct: 54 caatattcgccatgtttaaagcgatactaaaatttccttcactt 11
Database: est
Posted date: Jul 14, 2011 2:25 PM
Number of letters in database: 8,280,457
Number of sequences in database: 9484
Lambda K H
1.37 0.711 1.31
Gapped
Lambda K H
1.37 0.711 1.31
Matrix: blastn matrix:1 -3
Gap Penalties: Existence: 5, Extension: 2
Number of Sequences: 9484
Number of Hits to DB: 253,487
Number of extensions: 5129
Number of successful extensions: 5129
Number of sequences better than 1.0e-10: 1
Number of HSP's gapped: 5129
Number of HSP's successfully gapped: 1
Length of query: 1747
Length of database: 8,280,457
Length adjustment: 17
Effective length of query: 1730
Effective length of database: 8,119,229
Effective search space: 14046266170
Effective search space used: 14046266170
X1: 11 (21.8 bits)
X2: 15 (29.7 bits)
X3: 50 (99.1 bits)
S1: 12 (24.3 bits)
S2: 34 (67.9 bits)

